Cost Trends and Accessibility of Genome Sequencing Over Time

I still remember the shock of seeing a $15,000 invoice for sequencing a single human genome back in 2008. Our tiny four-person cancer research team had barely managed to scrape together enough grant money for two samples. The frustration was real—we knew the data could open doors to breakthroughs, but that price felt like a locked gate. Fast forward to today, and you can routinely sequence a whole genome for under $600. It’s not just about numbers dropping; it’s how those shifts rewired what we can actually do in both research and clinical care.
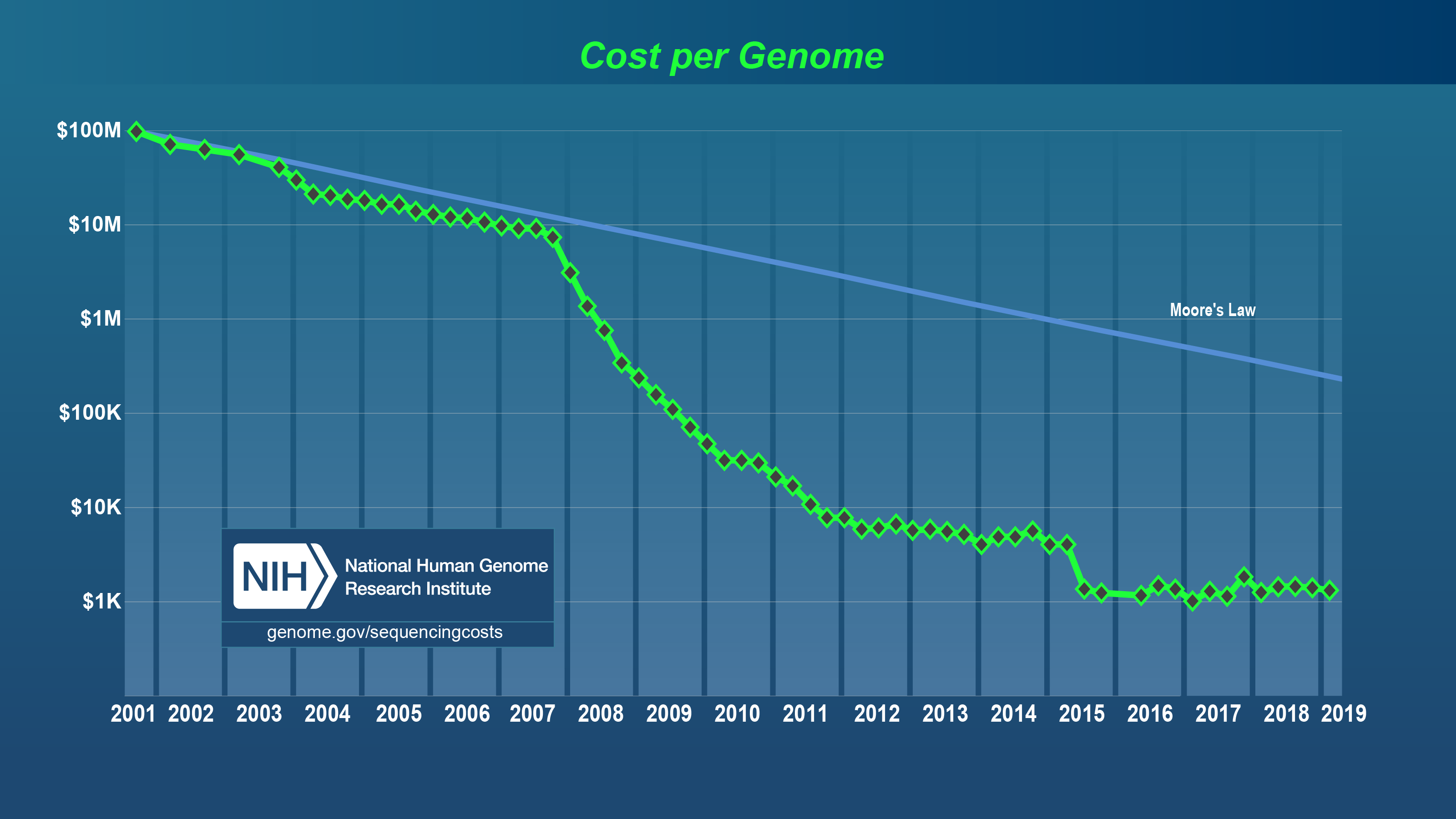
From $100 Million to Hundreds of Dollars: What You Don’t Hear Often
Most timelines jump from the Human Genome Project’s staggering $100 million price tag in 2001 straight to “next-generation sequencing slashed costs.” But that misses the messy middle. Back then, sequencing was a marathon slog using Sanger methods—picture technicians carefully running capillary electrophoresis machines and handling fluorescent dyes with zero margin for error. It wasn’t just pricey equipment; labor was intense, slow, and impossible to scale.
I once visited a sequencing core around 2003 where staff joked about “babysitting” machines overnight because any hiccup meant weeks lost. Meanwhile, computers struggled to process even megabytes of data fast enough. This bottleneck meant whole-genome sequencing was only within reach of government projects or pharma giants.
Next-Generation Sequencing (NGS): The Real Game Changer
Around 2007, when Illumina’s HiSeq systems landed in our lab, everything began shifting—though not overnight. Our first NGS run cost about $10,000 per genome—still steep but promising. The secret? Parallelization. Instead of decoding one DNA fragment at a time, these machines read millions simultaneously using bridge amplification and fluorescently tagged reversible terminators.
I remember staring at those first flow cells humming away as gigabases streamed out over days instead of months. Suddenly, we could dream bigger: our cohort exploded from 10 to 100 samples within two years as prices dropped below $2,000 per genome.
Why “Cheap” Sequencing Isn’t Always Cheap
Here’s something I wish I’d realized sooner: even when sequencing dips below $1,000 per genome, your project might still not be cheap or easy. Library prep kits alone often run near $200 per sample—and that’s before bioinformatics enters the picture.
Raw sequence data is just noise without skilled analysis pipelines. Early on, I spent weeks manually aligning reads because our computational tools couldn’t keep up—that frustration is very real if you don’t plan ahead.
If you’re on a lean budget—like when I consulted for startups developing personalized diagnostics—expect bioinformatics and interpretation to gobble up at least 40% of your total spend. Plus, turnaround times can be deceptive: some low-cost providers have queues stretching eight weeks or more. For clinical cases needing speed (think oncology), that delay can be fatal.
Real-World Example: Scaling Cancer Genomics Studies
During my postdoc at a mid-sized university lab in 2015, we faced a tough choice: sequence 50 tumor-normal pairs at around $2,500 each or switch to cheaper targeted panels with limited insight? We stuck with whole genomes because discovery beyond known mutations mattered.
By 2019, thanks to improved chemistry and competition (Oxford Nanopore shaking things up), costs dipped below $1,000 per genome. That allowed us to expand to over 600 patients in two years—a leap that directly led us to uncover rare variants linked to drug resistance no panel would have caught.
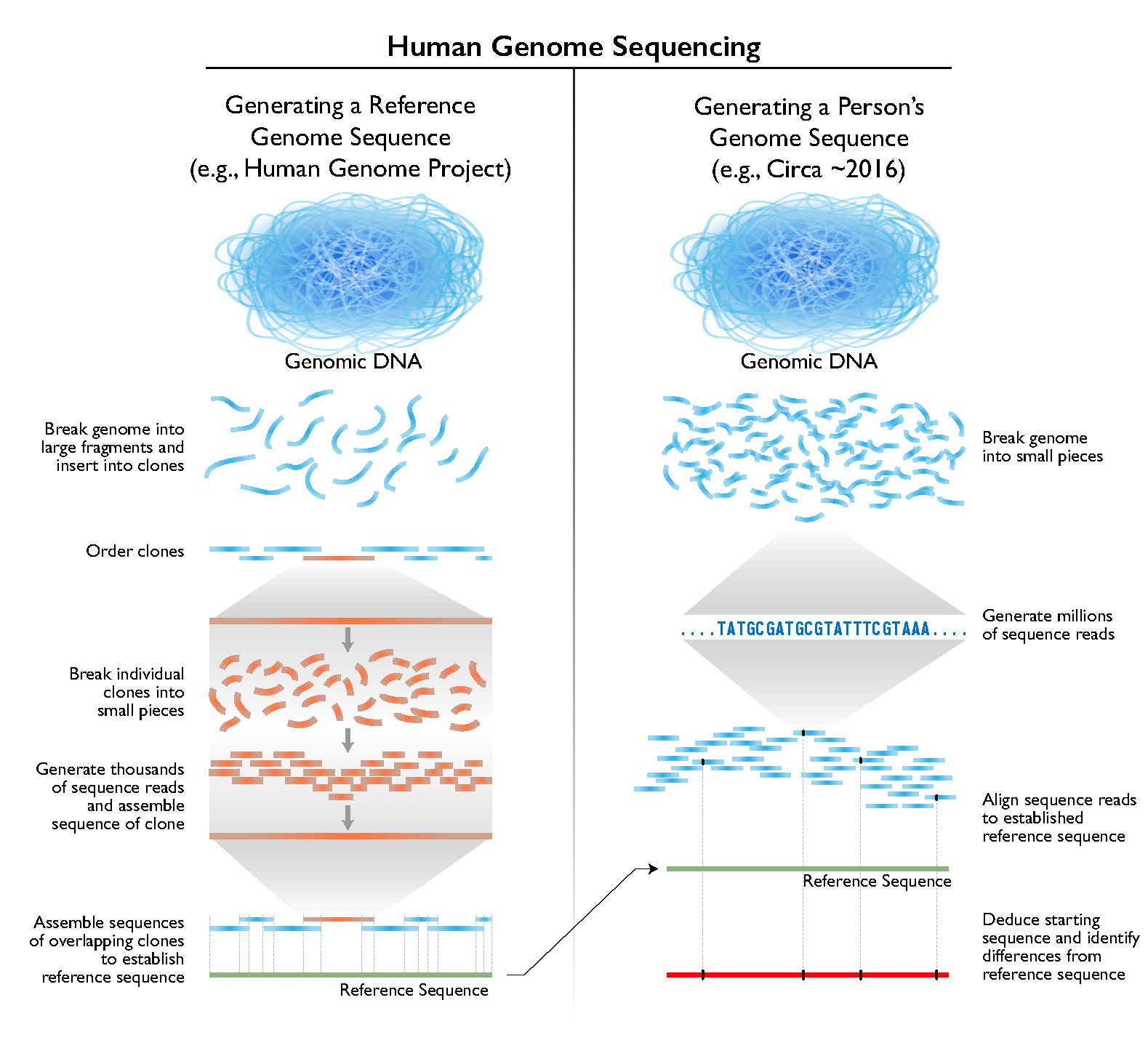
Accessibility Goes Beyond Price Tags
Lower sequencing prices don’t automatically mean global accessibility. In 2021, while consulting on a public health genomics project in Southeast Asia focused on infectious disease surveillance, we ran into hurdles beyond budget:
- Unreliable electricity tripped portable sequencers like Oxford Nanopore MinIONs
- Local regulations restricted sample export
- Skilled bioinformaticians were scarce
This taught me accessibility is a mosaic—not just technology cost but infrastructure and policy too. Programs like GA4GH are crucial here but progress varies wildly by region.
Lessons From the Trenches: How to Navigate Today’s Genome Sequencing Landscape
If you’re thinking about jumping into genome sequencing now, here are some battle-tested tips from my own ups and downs:
-
Don’t Trust Per-Genome Prices Alone: One provider might advertise $600 per genome but tack on $250 for library prep plus another $300 for bioinformatics analysis. I’ve chased “cheap” quotes that ballooned unexpectedly—ask for full breakdowns upfront.
-
Turnaround Time Matters: When my team needed rapid results for clinical trial enrollment in 2022, paying an extra $400 for expedited sequencing (under two weeks) beat waiting six weeks at half the cost every time.
-
Pilot Small Before Scaling: Running a test batch of 5–10 samples helped us identify prep variability and optimize pipelines before committing big bucks—it saved us tens of thousands later.
-
Use Hybrid Approaches: If budgets are tight but you want depth, combining targeted panels with low-pass whole-genome sequencing gave us broad coverage plus high-confidence variants at reduced cost.
-
Build Bioinformatics Partnerships Early: Raw data without solid analysis is worthless—partner with computational teams or use cloud platforms like Terra or DNAnexus right from the start rather than scrambling after sequencing completes.
Why Now Is Different — And Why Waiting Could Cost You More
Innovation isn’t slowing down—nanopore sequencers now fit in your pocket and deliver real-time DNA reads onsite. Last year in Uganda during an infectious disease genomics training trip, I saw MinIONs diagnosing viral outbreaks within hours—something unimaginable when I started this journey.
Waiting for prices to drop further risks missing windows where genomics can transform care or research today. My early mistakes taught me one thing clearly: thoughtful planning and choosing the right tech partner beats holding out for the “next big drop.”
Quick Key Takeaways
- Genome sequencing costs have plummeted—from ~$100 million in 2001 Human Genome Project days to under $600 now—but hidden costs remain significant
- Library prep ($150–$250/sample) and bioinformatics (often >40% total spend) require careful budgeting
- Turnaround times vary widely; faster results usually cost more but can be critical clinically
- Pilot projects save money by catching issues early
- Combining targeted panels with low-pass WGS can balance depth vs cost
- Accessibility depends on infrastructure & policy as much as price
- Start small; build partnerships early; pick providers who give transparent pricing
A Final Honest Thought
If you’re stepping into this world fresh: don’t let headline prices fool you into thinking it’s straightforward or cheap overnight. Technology choice, sample quality, bioinformatics capacity—they all drive costs and complexity more than most realize at first glance.
The revolution from $100 million per genome down to under $600 wasn’t just about tech—it was strategic patience mixed with constant adaptation over years (and lots of coffee-fueled late nights). Your best move? Start small, plan carefully, partner well—and get your data telling its story sooner rather than later.
If you want specific provider recommendations or help designing workflows tailored exactly to your budget and goals—reach out anytime! Having stumbled down this path more than once means I can help you dodge common traps and make every dollar count toward real discovery.
The genomic frontier isn’t tomorrow anymore—it’s happening now—and it’s waiting for you too.
If you're curious about providers: Illumina remains dominant for high-throughput short-read sequencing; Oxford Nanopore offers affordable portable options great for rapid field work; PacBio shines when long reads matter most. For cloud analysis platforms: Terra (by Broad Institute) and DNAnexus are reliable starting points with user-friendly pipelines ready-made for many applications.
And yes—I once accidentally dropped a tube containing precious library prep reagents onto the floor right before loading them into the sequencer... talk about heart-stopping moments! But hey—that’s part of the game we all learn from along the way.



