Histone Modification: How It Controls Gene Regulation Simply Explained

When I first stumbled into the world of histone modifications, it honestly felt like cracking a secret code written between the lines of DNA. It’s not about changing the genetic letters themselves, but more like whispering instructions: “Turn me up,” or “Shush now.” Most beginner guides make it sound like acetylation or methylation are simple on/off switches for genes, but after years spent chasing these tiny chemical marks in the lab, I can tell you — it’s way messier and way cooler than that.
Here’s what took me a while to grasp: histone modifications aren’t just digital toggles flipping gene expression on or off. They’re part of a delicate dance where timing, combinations, and context matter so much more than any single mark alone. If you want to move beyond textbook summaries and actually harness these modifications in your experiments, buckle up—I’ll share some hard-won lessons from my own research journey.
Histone Modifications: More Than Just DNA Wrapping
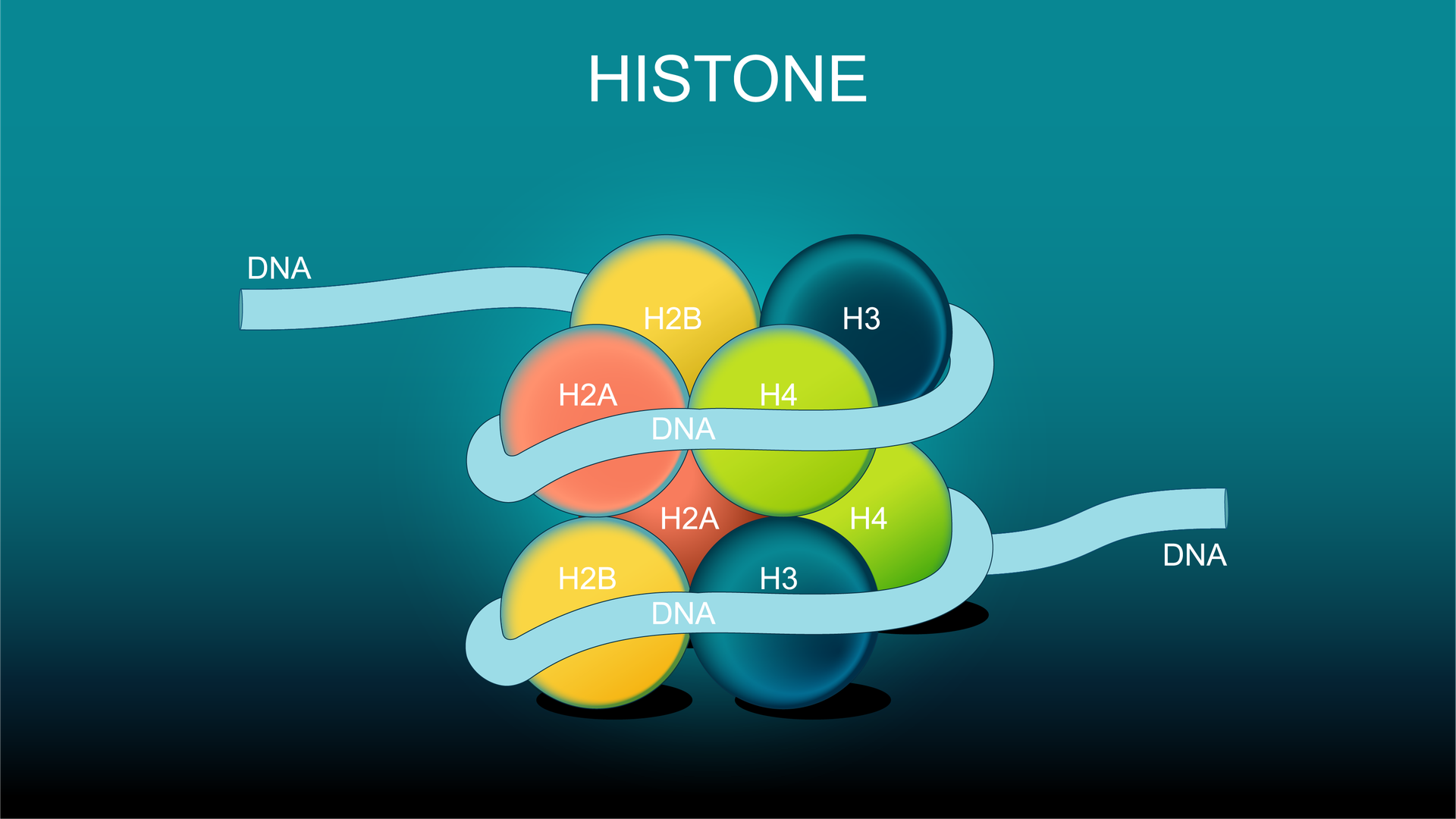
Picture DNA as a long thread wrapped around spools called histones. How tightly or loosely that thread winds controls whether certain genes are visible or hidden away. Now, the magic happens when small chemical groups—like acetyl (–COCH₃) or methyl (–CH₃)—attach to specific amino acids on the histone tails sticking out from those spools.
You’ve probably read things like: “Acetylation relaxes chromatin; methylation can either activate or repress gene expression.” But from months of ChIP-seq experiments, here’s my reality check: that’s an oversimplification.
For example: H3K4me3 is often labeled an “active” mark—but in some cells I studied, regions with H3K4me3 weren’t fully active; they were poised, waiting silently for the right signal to spring into action. The cell’s environment, other epigenetic marks nearby, even metabolic state—all these factors decide if poised means go or stay quiet.
So don’t just memorize “H3K27me3 = silent” without remembering context matters deeply.
Acetylation and Methylation: The Dynamic Duo With Surprises
Everyone says acetylation neutralizes positive charges on histones to open up DNA for transcription. That’s true! But here’s something many don’t mention: the enzymes responsible—histone acetyltransferases (HATs) and deacetylases (HDACs)—never work solo. They get recruited by “reader” proteins that interpret these modifications and decide what happens next.
I’ll never forget my early assumption that adding acetyl groups always cranks up transcription. In one biotech startup I consulted for, we saw hyperacetylation correlate with gene repression—not activation—a total head-scratcher at first! After digging in, we found this was due to crosstalk with other histone marks and recruitment of repressive complexes that bind specifically to those acetylated tails in certain cellular contexts.
Methylation adds even more complexity. Take mono-, di-, and tri-methylation on lysines—each can mean wildly different things depending on which lysine is modified and what stage of development or disease you’re looking at.
Real-World Example: The HOX Genes’ Epigenetic Dance
One of my favorite examples comes from studying HOX genes—the masters of body patterning during embryonic development. Their regulation perfectly showcases how histone modification dynamics work in real time.
In early embryos, Polycomb group proteins slap down heavy H3K27me3 marks across HOX clusters like locked vaults keeping them silent. As development proceeds, those repressive marks get selectively erased, while activating marks like H3K4me3 appear precisely where and when needed—unlocking genes step-by-step.
I spent nearly two months trying to replicate this choreography in cultured stem cells using ChIP-seq combined with RNA-seq data integration. At first? Total failure—the timing didn’t match up at all. Then I learned something crucial: epigenetic changes often precede transcriptional activation by many hours or even days. When we stretched our time-course experiments over 72 hours instead of just 24, we finally saw the stepwise eviction of repressive marks followed almost 48 hours later by gene expression turning on.

Patience was key here—and it’s something every newcomer should brace for!
Insider Tips From My Lab Notebook
-
Validate your antibodies rigorously
I wasted weeks chasing false signals because my anti-H3K27me3 antibody cross-reacted with other methylated residues. A quick peptide competition assay saved me from going down that rabbit hole further—don’t skip this step! -
Beware bulk averaging in ChIP-seq
If you’re working with complex tissues made of mixed cell types (hello brain samples), your signals will blur key cell-specific differences. Whenever possible, pair ChIP-seq with single-cell ATAC-seq or CUT&Tag methods—they zoom in on cellular heterogeneity so you don’t miss subtle but important changes. -
Cross-validate histone marks with RNA expression and functional assays
Seeing a peak isn’t enough! Too many projects stop there without confirming if that gene’s actually changing its expression—or worse, assuming correlation means causation right away. -
Don’t overlook less famous modifications
Ubiquitination or phosphorylation on histones might seem like footnotes compared to acetylation/methylation—but they often fine-tune regulation in ways those big players can’t explain alone. For example, H2B ubiquitination plays a crucial role in facilitating subsequent methylations during active transcription—something worth digging into when things don’t add up!
A Counterintuitive Lesson: Focus Beats Frenzy
Early on, I tried mapping every possible histone modification across every gene under the sun thinking it would untangle all gene regulation mysteries instantly. Nope—it just created a confusing combinatorial explosion!
Our breakthrough came when we focused narrowly on just two key modifications—H3K27ac and H3K4me3—in a cancer cell line over six months. By integrating these with RNA-seq data at multiple time points after drug treatment, we pinpointed core regulatory shifts driving response versus resistance—no guesswork needed.
So here’s my advice: pick your battles wisely and dig deep into a few well-chosen marks combined with multi-omics data rather than spreading yourself too thin chasing every mark out there.
What I’d Tell My Younger Self (or You)
If you take away only one thing from this whole story: never treat histone modifications as simple binary switches — they’re part of a complex conversation influenced by cell type, developmental stage, environment—and most importantly—the interplay between multiple epigenetic mechanisms working together.
Get comfortable living with ambiguity at first—it took me years before I could confidently say “this mark means this” without caveats hanging over my head.
But once you embrace that complexity? You unlock a profoundly rich layer of biological regulation hidden within chromatin—a perspective that shifts your entire understanding of gene expression beyond just DNA sequence alone.
Quick-Start Checklist for Your First Chromatin Experiment
- ✅ Validate antibodies by peptide competition assays before trusting results
- ✅ Use time-course designs; epigenetic changes often precede transcription shifts
- ✅ Complement ChIP-seq with RNA expression data (RNA-seq or qPCR)
- ✅ Consider single-cell techniques if working with heterogeneous samples
- ✅ Start focused: pick 1–2 key modifications relevant to your system rather than all at once
- ✅ Don’t ignore less common modifications (ubiquitination/phosphorylation) — keep them in mind if results puzzle you
Got stuck navigating ENCODE datasets? Need tips designing your first ChIP-qPCR experiment? Feel free to ask—I’ve been down those frustration-filled roads myself and can share shortcuts that saved me months of headaches!
Remember—science is messy and slow sometimes; patience paired with curiosity will get you farther than rushing through protocols ever will.
Happy chromatin hunting!



