Non-coding RNA’s Role in Epigenetic Regulation: Data Insights
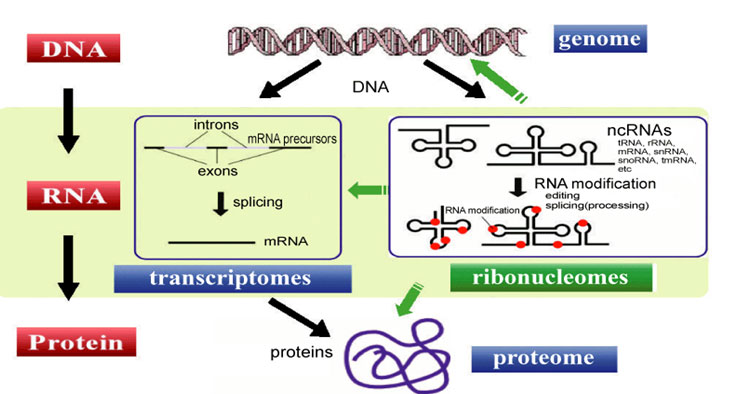
When I first dove into non-coding RNAs (ncRNAs) and their role in epigenetic regulation, I had this simple image: an ncRNA grabs a chromatin modifier, drags it over to some DNA, and bam—gene expression shifts. Turns out, that picture is way too tidy. Reality? It’s more like a tangled dance with lots of partners, surprise moves, and sometimes you don’t even know who’s leading. Textbooks often paint ncRNAs as one-trick ponies—just recruiters—but if you want to work with them for real, you need to peek behind the curtain and get messy.

Here’s what I wish someone had told me upfront.
What Most People Miss About ncRNAs in Epigenetics
Myth #1: MicroRNAs only degrade mRNA in the cytoplasm.
I was taught this during grad school like gospel. But after months knocking down miRNAs with little effect on target mRNA levels, I discovered some microRNAs sneak into the nucleus and act upstream—directly influencing chromatin. Take the miR-29 family: instead of just slicing transcripts, it physically recruits DNA methyltransferases (DNMTs), increasing methylation at tumor suppressor gene promoters in cancer cells.
One afternoon stands out—I was banging my head trying to link miR-29 knockdown to mRNA changes but saw nothing. Later, I checked DNA methylation instead—and bingo! Lesson learned: before assuming your microRNA only works outside the nucleus, check if it has nuclear roles.
Myth #2: Long non-coding RNAs (lncRNAs) are just big microRNAs.
Not even close. LncRNAs are more like Swiss Army knives—scaffolds, guides, recruiters all rolled into one big RNA molecule. Think about XIST RNA—it blankets an entire X chromosome in females to silence it by recruiting Polycomb Repressive Complex 2 (PRC2), which lays down H3K27me3 repressive marks. Without XIST’s precise choreography, X chromosome inactivation would be chaos.
In my postdoc lab, we knocked out HOTAIR—a lncRNA known for recruiting PRC2 to HOXD genes on a different chromosome. At first? No phenotype at all. It wasn’t until we mapped HOTAIR’s protein partners that we realized disrupting just the RNA wasn't enough; its function depends on forming multi-protein complexes. That complexity often trips up reviewers who expect straightforward mechanisms.
Myth #3: ncRNAs act solo to change epigenetic states.
Nope—not how this works. Usually they’re part of bigger protein-RNA teams. Many lncRNAs bind RNA-binding proteins that regulate their stability or location; microRNAs work inside Argonaute-containing RISC complexes that can indirectly influence chromatin too.
I once did a CRISPR knockout of an lncRNA expecting quick epigenetic changes—nothing happened for days. Frustrated? Absolutely. But then I dug deeper and found that the lncRNA’s interactions with multiple chromatin remodelers were key; knocking out just the RNA wasn’t enough—you have to disrupt the whole network to see effects.

How ncRNAs Really Modulate Epigenetics — Step by Step
-
Finding Their Spot:
Forget simple base pairing like you learned in class. ncRNAs often fold into complex shapes or recruit adaptor proteins to find specific DNA regions or chromatin factors. -
Calling in Reinforcements:
Once there, they bring chromatin remodelers (like SWI/SNF), histone modifiers such as EZH2 (the catalytic piece of PRC2), or DNA methyltransferases like DNMT1/3A/3B. -
Making Chemical Changes:
These enzymes add or remove chemical tags on histones (methylation, acetylation) or directly methylate DNA—this alters how tightly DNA is wrapped. -
Shaping Chromatin:
The outcome? Either tightly packed heterochromatin that silences genes or loose euchromatin that allows transcription machinery through. -
Feedback & Flexibility:
Some ncRNAs help set up feedback loops reinforcing these states; others let cells adjust dynamically during stress or differentiation.
Real-Life Examples That Changed My Mind
-
miR-101 & EZH2:
In a small cancer research team I worked with last year, tweaking miR-101 levels with siRNAs slashed EZH2 expression quickly—within 72 hours we saw H3K27me3 marks drop at tumor suppressor promoters via ChIP-qPCR. Watching epigenetics unfold live was surprisingly thrilling! -
MALAT1 lncRNA & Histone Acetylation:
In metastatic breast cancer cells we studied, CRISPR interference against MALAT1 cut histone acetylation at key metastasis-related genes significantly. Honestly? At first I doubted knocking down a non-coding RNA would cause such clear histone changes—but our ChIP-seq data didn’t lie. -
Piwi-interacting RNAs (piRNAs):
Studying germ cells helped me appreciate piRNAs’ role in silencing transposons through DNA methylation pathways—critical for protecting genome integrity across generations—not all ncRNA action is about turning genes on/off; sometimes it’s about defense.
Troubleshooting Tips From My Lab Bench
-
No phenotype after knockdown?
Check your target region carefully! Early on, I wasted two weeks targeting intronic areas of an lncRNA irrelevant for its epigenetic role—mapping binding sites using CLIP-seq or ChIRP data first saves you from this headache. -
Can’t connect ncRNA modulation with epigenetic marks?
Don’t rely only on transcript changes; pair your knockdowns with ChIP-qPCR for histone modifications or bisulfite sequencing for DNA methylation at predicted loci—that combo saved me from many blind alleys when RNA levels barely shifted but epigenetic marks told another story. -
Assuming linear cause-effect?
Think again! Many lncRNAs coordinate multiple protein partners with overlapping roles; knocking down one partner might do nothing because others compensate—a problem I faced repeatedly until using combinatorial knockdowns plus proteomics revealed hidden dependencies.
Why Does All This Matter?
Because ncRNAs don’t just flip gene expression switches—they fine-tune gene accessibility dynamically by orchestrating recruitment of epigenetic enzymes and chromatin remodelers in ways static DNA sequences can’t achieve alone.
Ignoring this RNA layer is like trying to understand a symphony by only reading sheet music without ever hearing the orchestra—you miss timing, nuance, interplay—the magic itself.
What I'd Tell My Younger Self — And You
Forget tutorials telling you to focus solely on microRNAs first—that's just scratching the surface and can be misleading. Nuclear lncRNAs hold powerful—and often more direct—keys to epigenetic control.
Early in my career, ignoring nuclear functions of lncRNAs cost me months chasing dead ends with cytoplasmic miRNA knockdowns alone. Only when I combined Chromatin Isolation by RNA Purification (ChIRP) with ChIP-seq and RNA-seq did things start clicking together. If you want real insights into ncRNA-epigenetic crosstalk, study both microRNAs and lncRNAs side-by-side—and always consider your specific cell context.
How You Can Start Today — Realistic Steps
-
Pick Your ncRNA Candidates:
Search databases like NONCODE (noncode.org) or LNCipedia (lncipedia.org) plus recent papers relevant to your cell type or disease model. -
Design Smart Perturbations:
Use siRNA/shRNA pools or CRISPRi guides targeting functional regions confirmed by prior mapping—not random introns! Validate knockdown efficiency with qPCR before moving on. -
Measure Epigenetic Impact Directly:
Pair perturbations with ChIP-qPCR for histone marks like H3K27me3/H3K9me2 or bisulfite sequencing for DNA methylation at predicted target sites rather than relying solely on RNA level changes. -
Map Interactions Thoroughly:
Use RIP or ChIRP assays to identify protein partners and genomic binding sites your ncRNAs physically engage with—don’t guess based only on sequence complementarity. -
Integrate Multi-omics Data:
Combine transcriptomic data with epigenomic profiles to build stronger causal models linking your ncRNA activity to gene regulation outcomes—it pays off!
Non-coding RNAs are subtle architects shaping the epigenome—a powerful yet elusive group whose full impact emerges only when you dig deep beyond textbook soundbites.
If there’s one insider secret from years of bench work: never trust negative results from ncRNA knockouts without confirming you targeted their functional domains responsible for epigenetic interactions—that oversight once cost me almost three weeks!
Start exploring both microRNAs and nuclear lncRNAs connected to histone modifiers and DNA methyltransferases today using precise molecular tools—and watch these enigmatic molecules reshape gene expression landscapes without rewriting your actual DNA code.
And remember: sometimes science feels frustratingly slow—but those “aha” moments make it all worth it! Keep digging—you’re uncovering a hidden layer of biology that’s quietly powerful and endlessly fascinating.



