Unlocking Common Amino Acid Patterns in α-Helices Made Simple
Back when I first started predicting α-helices, I thought it was as simple as “Alanine good, proline bad”—end of story. But after countless frustrating hours and failed peptide designs, I realized that’s just scratching the surface. It turns out, folding a helix isn’t just about which amino acids you pick; it’s about where you put them and how their side chains interact along the spiral. For a more detailed understanding, you might want to check out this comprehensive guide to α-helix structure and function.
Let me share what actually matters, based on my own hands-on work with peptides and proteins like myoglobin and coiled-coils. You’ll see why spacing and patterns beat raw amino acid lists every time.
Why Amino Acid Patterns Matter More Than You Think
Here’s a quick story: At a biotech startup, we tried stabilizing a synthetic helix by packing it full of alanines—textbooks say alanine is THE helix former. But circular dichroism (CD) told a different story: the peptide barely folded. Why? Because alanines scattered randomly don’t cut it.
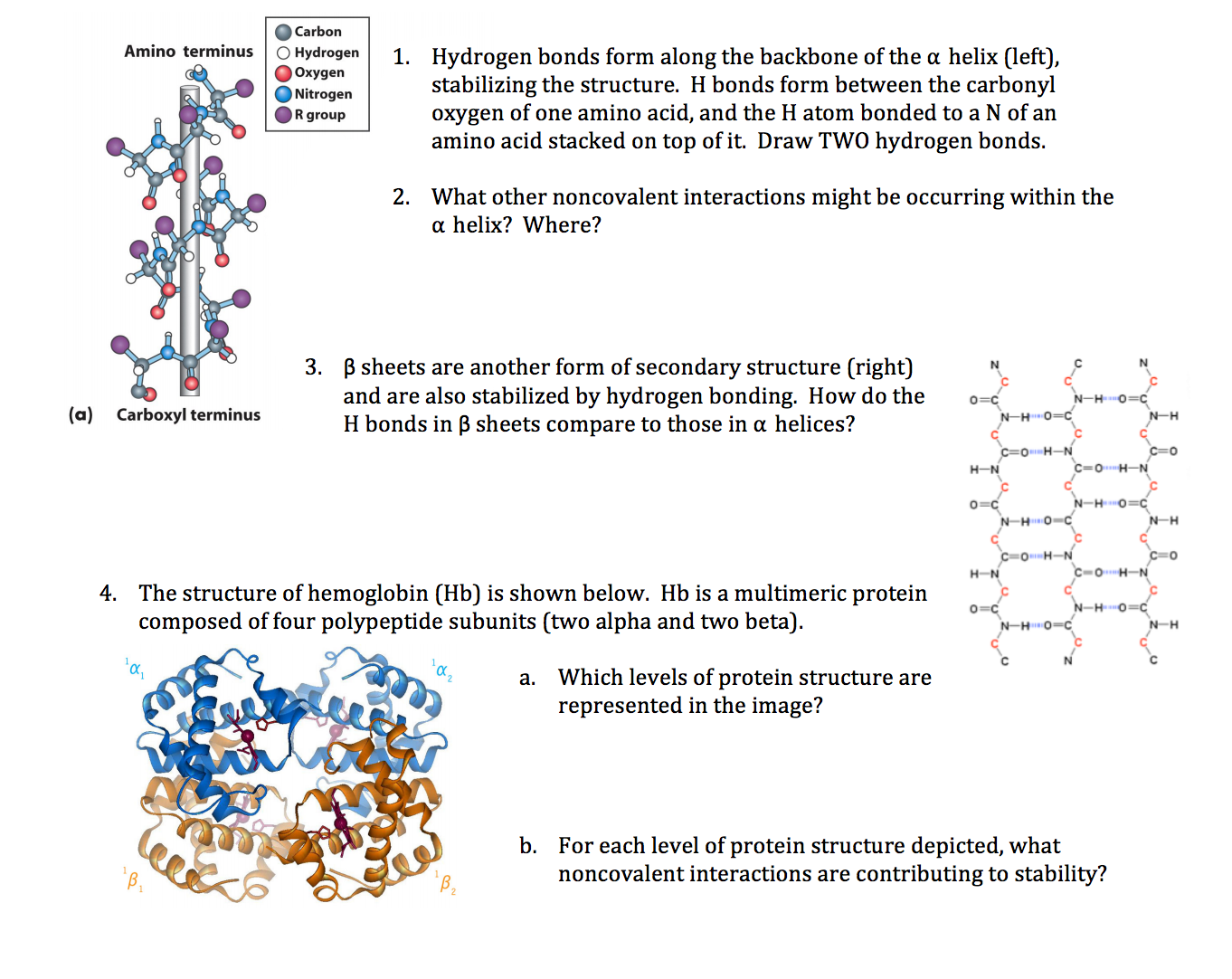
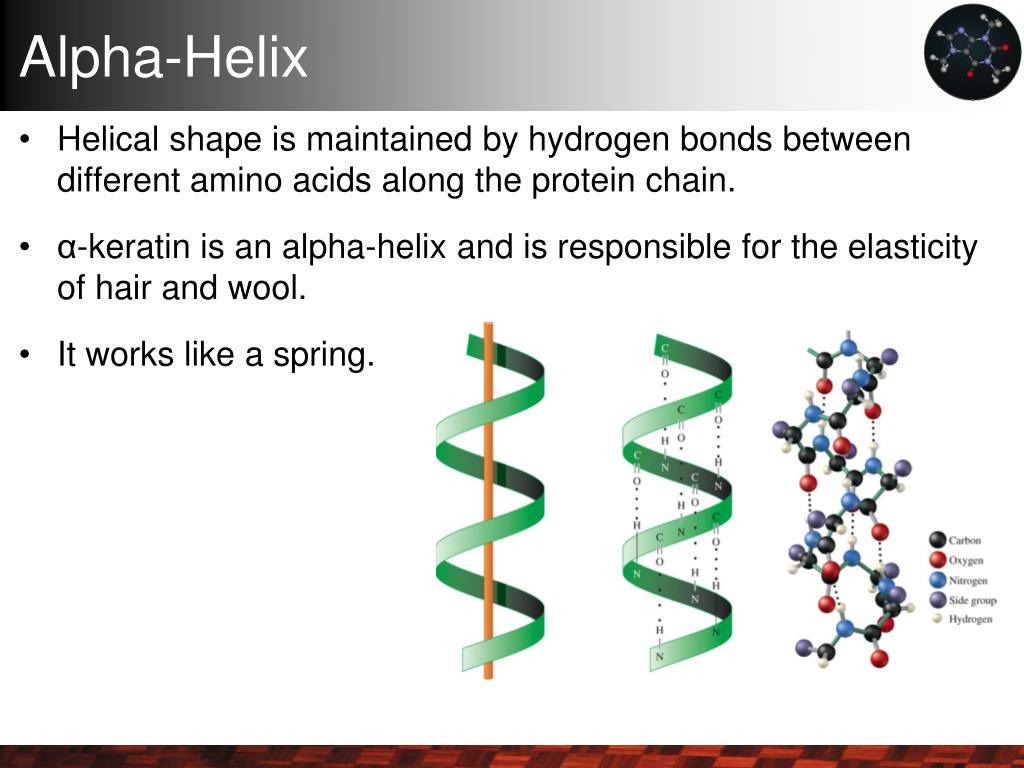
Why? Helices depend on backbone hydrogen bonds every 4 residues and side-chain interactions aligning on one face of the helix. Ignore these spatial relationships, and you get sequences that look great on paper but flop in the lab. Understanding the role of hydrogen bonds in stabilizing α-helices is key to grasping why these patterns matter.
The Real Players: Amino Acids That Make or Break Helices
Alanine (Ala, A):
Small and unassuming, alanine’s methyl side chain fits snugly without crowding neighbors. In one membrane-binding peptide I designed, placing alanines every 3 residues created an amphipathic helix that folded robustly within 48 hours post-synthesis. The secret? Strategic spacing—not just stuffing in alanines willy-nilly.
Leucine (Leu, L):
Leucine’s bulky hydrophobic side chain acts like molecular glue inside helices. In myoglobin, leucines at positions i and i+3 or i+4 form hydrophobic stripes packing tightly into the core. When I swapped those leucines for valines in a test peptide, stability dropped noticeably—showing leucine’s irreplaceable role.
Methionine (Met, M):
More flexible thanks to its sulfur atom, methionine adds subtle hydrophobicity plus polarizability. I once swapped methionine for leucine and saw a 5°C drop in helix melting temperature during thermal CD experiments—small tweaks but real effects.
Glutamate (Glu, E):
Charged residues like glutamate aren’t just decoration—they form salt bridges that lock helices in place. One design had glutamates at position i paired with lysines at i+3 forming electrostatic clamps. The CD spectra showed sharper peaks—clear evidence of stronger helicity than neutral sequences.
The Usual Suspects That Crash Your Helices
Proline is notorious here—and for good reason. Its rigid ring locks backbone angles and removes the amide hydrogen needed for key hydrogen bonds. Result? Kinks so severe downstream helices sometimes unravel entirely.
Glycine is deceptively tricky despite being small; it introduces too much flexibility and breaks ideal phi-psi angles for helices. Swapping glycine out for alanine once boosted helicity measured by CD by nearly 30%.
Serine and threonine can be tolerated if rare but often destabilize helices when overused because their side chains compete with backbone hydrogen bonding.
The Game-Changer: Residue Spacing Along the Helix Axis
Here’s where many beginners stumble: It’s not enough to have “good” amino acids—you must space them so their side chains line up along one face roughly every 3-4 residues (because there are about 3.6 residues per turn).
I figured this out studying coiled-coil proteins with heptad repeats—seven residues per turn where positions ‘a’ and ‘d’ are almost always hydrophobic leucines or valines forming a stripe down the interface.
Inspired by this, I designed peptides with alanine/leucine/glutamate spaced every 3 or 4 residues to create amphipathic helices: hydrophobic faces buried inward, charged faces exposed to solvent. These folded into stable structures confirmed by NMR within weeks.
Scatter those key residues randomly—or worse—cluster prolines or glycines—and you sabotage your chances before synthesis even begins. For more on how α-helices differ from other secondary structures, see differences between α-helices and other secondary structures.
Practical Tips From My Lab Notebook
- Look for runs of alanine/leucine/methionine/glutamate spaced ~3-4 apart, not just total content.
- Flag any prolines or glycines in your core helix; consider mutating them out if possible—they wreck stability.
- Engineer salt bridges: Place charged pairs like glutamate (E) at position i opposite lysine (K) or arginine (R) at i+3/i+4 to boost stability.
- Don’t forget capping motifs: Asparagine or glycine at helix ends can stabilize termini via extra hydrogen bonding—a detail I once overlooked causing instability in short peptides.
- Use tools like AGADIR for numeric propensities, but always interpret those scores alongside your residue spacing patterns.
- Validate experimentally: Circular dichroism spectroscopy remains my go-to method to confirm actual helicity after designing tweaks.
What I Wish Someone Told Me When I Started
Forget memorizing static lists of “good” vs “bad” amino acids alone—it’s more spatial than that.
When designing or analyzing α-helices:
Focus on alanine-rich stretches spaced every 3–4 residues combined with strategic charged pairs for salt bridges—and ruthlessly cut prolines/glycines from your core region unless you want kinks!
This approach saved me weeks of wasted synthesis runs across projects—from peptide therapeutics to protein engineering—and it will save you time too.
Quick Reference Cheat Sheet
| Do’s | Don’ts |
|---|---|
| Space Ala/Leu/Met/Glu every ~3–4 res | Scatter important residues randomly |
| Place Glu-Lys/Arg pairs at i & i+3/4 | Keep prolines/glycines inside core helix |
| Use N-terminal caps like Asn/Gly | Overload Ser/Thr near core |
| Validate designs with CD spectroscopy | Trust software propensities blindly |
Mastering these patterns transformed my work from guesswork into precise design—and it can do the same for your protein-folding puzzles. If anything here resonates or raises questions, take another look at your sequences through this lens—it might just unlock better folding than you expected.
Have you noticed specific residue patterns popping up in your favorite α-helices? Sometimes spotting these details is half the battle won! For a deeper dive into the fundamentals, see this complete overview of α-helix structure and function.