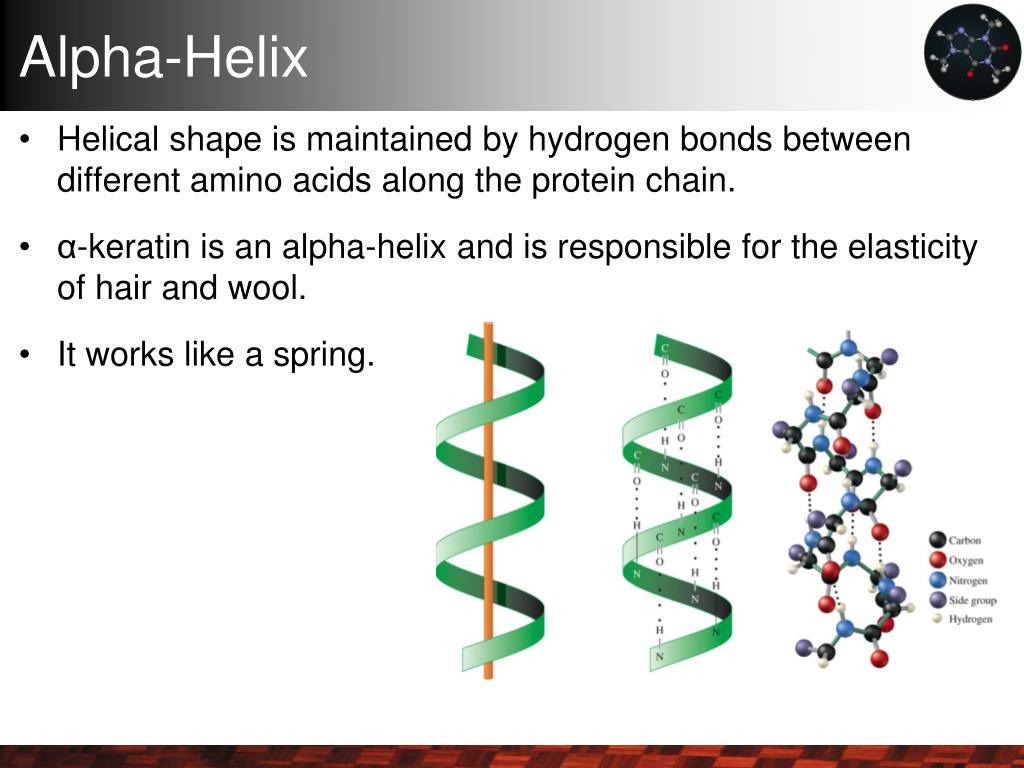
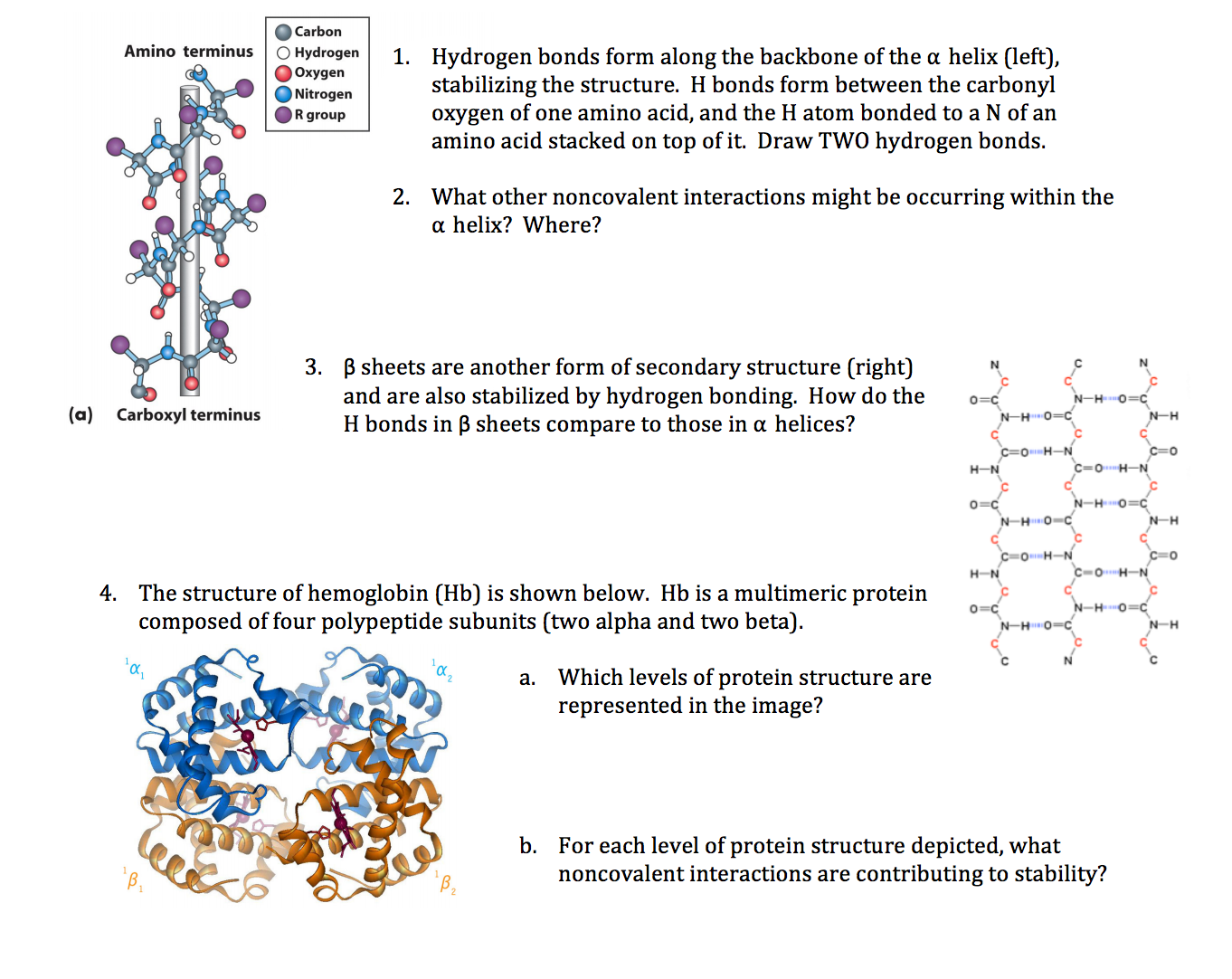
Unlocking the Power: Functional Importance of α-Helices in Membranes

When I first dove into studying membrane proteins, I was honestly overwhelmed by how much α-helices were treated like boring, rigid scaffolds. It took me months—yes, MONTHS—of chasing confusing mutagenesis results before I realized something crucial: α-helices aren’t just passive building blocks. They’re dynamic, finely tuned players that respond to their lipid environment in ways that can make or break protein function. If you want to deepen your understanding, check out this comprehensive guide to α-helix structure and function.
So here’s the real deal, from someone who’s been in the trenches—not theory or textbook fluff: α-helices dominate membrane proteins because they strike a delicate balance between holding structure and adapting to the oily, harsh membrane. Understanding them means focusing on that balance rather than drowning in overcomplicated models.
Why Do Membranes Love α-Helices So Much?
Membranes are tough neighborhoods for proteins—oily, hydrophobic, and starved of water. That’s why α-helices are the go-to secondary structure for membrane-spanning segments:
-
Picture a “helical wheel” — it’s like the helix is wrapped in a ring of side chains pointing outward. These side chains are mostly hydrophobic (think leucine, isoleucine), perfectly matching the fatty acid tails of the membrane. This snug fit isn’t optional; it’s essential. β-sheets just don’t cut it here because they need water molecules to stabilize their backbone.
-
Inside membranes, there’s no water to shield those backbone hydrogen bonds (the amides and carbonyls). The α-helix cleverly solves this by forming an internal hydrogen bond network that’s strong enough to keep shape but flexible enough to bend or kink when needed. For a deeper dive into how these bonds stabilize helices, see the role of hydrogen bonds in stabilizing α-helices.
I still remember one bacterial transporter I studied where a small kink in a TM helix — caused naturally by a proline residue — acted like a gatekeeper for ions. Without that bend, the channel didn’t open properly. So these kinks aren’t defects; they’re functional switches.
For a more detailed explanation of α-helix roles and dynamics, see the complete overview of α-helix structure and function.
More Than Just Stability: What Else Can Helices Do?
If you think helices only hold things together, think again:
-
Channel Gates and Selectivity: In ion channels I worked on, bundles of helices form tiny pores. Even slight tweaks in how these helices pack—like swapping one leucine for proline—dramatically changed which ions passed through and how often channels opened.
-
Signal Transmission: GPCRs are classic examples where helix rearrangements inside membranes transmit signals. When I introduced mutations that disrupted helix shape but left membrane insertion untouched, receptor activation changed drastically. The takeaway? Geometry matters as much as presence.
-
Protein Partnerships: In a multi-protein complex I helped with, certain helices served as “Velcro strips,” sticking proteins tightly together. Mutating just one residue on these interfaces broke assembly without disrupting folding—a subtle but critical role.
-
Lipid Sensing: Some helices even “read” their lipid surroundings. Changing hydrophobic residues facing lipids to polar ones shifted where the protein localized within membrane microdomains—and its activity changed accordingly. At first glance, this was baffling until I realized those helices interact directly with specific lipids.
How I Learned to Study Helices Without Spinning My Wheels
I wasted too many weeks throwing random mutations around before developing this simple—but powerful—strategy:
-
Confidently Identify Your Helices
Don’t just trust TMHMM or Phobius blindly—combine predictions with any structural data (X-ray or cryo-EM). If you can, perform limited proteolysis or antibody accessibility assays for experimental confirmation. For more on how α-helices differ from other secondary structures, check out differences between α-helices and other secondary structures. -
Mutate Smart
Focus on 2–3 strategic residues per helix:- Replace hydrophobic side chains facing lipids (like Leu or Ile) with charged residues (Glu or Lys) to disrupt interaction sharply.
- Introduce a proline smack in the middle to create a controlled kink—you’ll see clear functional effects sometimes without losing folding altogether.
- Target conserved glycines involved in helical packing; these tiny residues play big roles in tight helix packing.
-
Check Secondary Structure Directly
Circular Dichroism (CD) spectroscopy became my go-to for monitoring helical content changes post-mutation—it gives you direct evidence whether your mutation distorts the helix or not. No CD? Environment-sensitive dyes like ANS can give indirect clues about folding states. -
Confirm Membrane Insertion
Protease protection assays are surprisingly straightforward and reliable: if your protein loops are protected from digestion inside cells, you know insertion happened right. Surface biotinylation is another good complementary approach for topology verification. -
Link Structure to Function
Don’t stop at folding/insertion! Measure what really counts: ion flux for channels, substrate uptake for transporters, ligand binding or downstream signaling for receptors. Sometimes insertion is fine but function tanks—and that tells you helices do more than basic architecture. -
Use Molecular Dynamics With Caution
I found tools like GROMACS very helpful—but only after having solid experimental data first! Simulations can predict if mutations destabilize helix-lipid interactions but relying on them prematurely wastes time.
Learning From My Worst Mistakes
Here’s an honest confession: once I swapped a leucine for glutamate expecting just local disruption in a bacterial transporter TM helix—and lost all activity instead! Initially blaming charge repulsion was naive; CD showed partial unfolding too. Lesson learned: breaking backbone hydrogen bonds inside membranes is far worse than side chain chemistry alone suggests.
In another case with GPCRs, adding proline didn’t affect folding or insertion but completely screwed up downstream signaling patterns—classic example of how standard stability tests miss critical functional geometry changes.
These experiences hammered home that each helix has its own context-dependent roles intertwined with local lipids and protein partners—you can’t treat them all alike.
The Single Most Important Insight I’d Pass On
If you’re new here: don’t rush into massive modeling projects or shotgun mutagenesis screens before designing targeted mutations based on solid structural data—and always pair multiple orthogonal assays (folding + insertion + function). This triangulation cuts through noise and reveals what helices truly do inside membranes.
Ready To Start? Here Are Practical Steps You Can Take Today
- Run TMHMM on your sequence and mark predicted helices confidently—but verify with any experimental data you can find.
- Pick 2–3 hydrophobic residues per helix facing lipids (e.g., Leu→Glu or Ile→Lys) for targeted mutations.
- If possible, purify mutant proteins and check their helical content using CD spectroscopy.
- Set up protease protection assays on cells expressing your mutants.
- Design functional assays tailored to your protein’s role—ion flux for channels; ligand binding/signaling for receptors.
- If comfortable with simulations, run short MD trajectories focusing on mutated helices embedded in lipid bilayers—but only as support after experiments!
This approach saved me months of frustration and clarified how α-helices dictate not only structure but subtle functional outcomes in membranes.
A Parting Thought
Understanding α-helices isn’t about mastering complicated computational models—it’s about observing carefully what happens when you tweak these structures just so…and then listening closely to what your experiments tell you next. It’s messy sometimes; it takes patience; but real insight lies there, far from generic advice or oversimplified dogma.
If I could share one piece of advice from my journey: treat every helix as its own little world shaped by both protein neighbors and lipids—and never underestimate their power beyond mere scaffolding.
If you'd like some quick references:
- TMHMM Server – reliable transmembrane prediction
- Circular Dichroism Spectroscopy basics – intro article & protocols
- Protease Protection Assays overview – step-by-step guide
Feel free to ask if you'd want me to suggest specific protocols or troubleshoot interpretation down the road—I’ve been there more times than I care to admit!
Does this resonate? What challenges have you faced with α-helices so far? Sometimes sharing those ‘failures’ is where real progress begins...